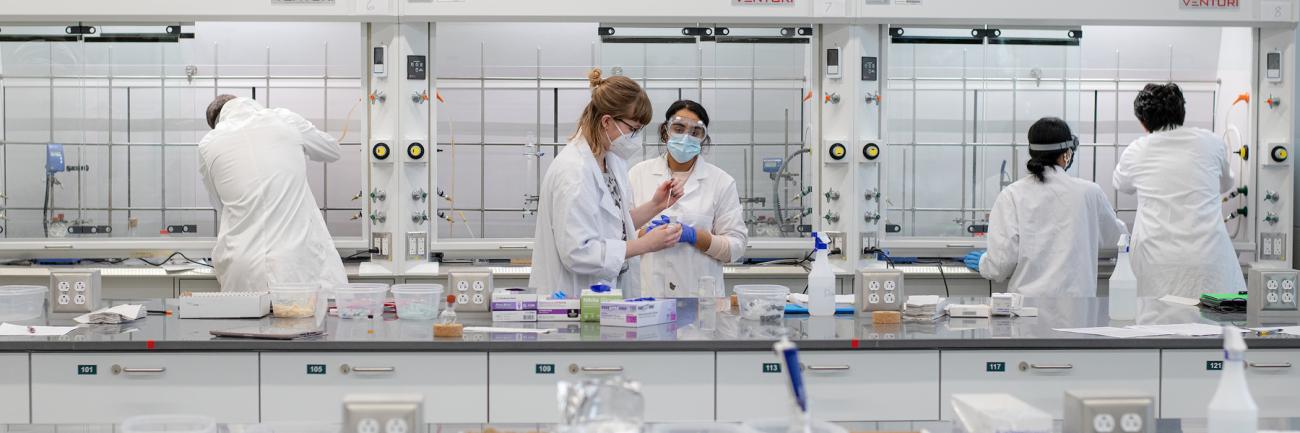
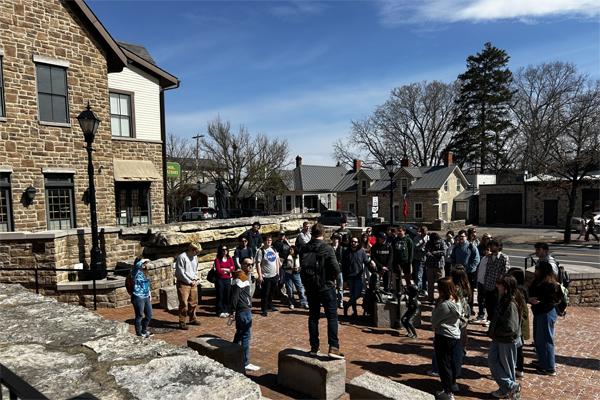
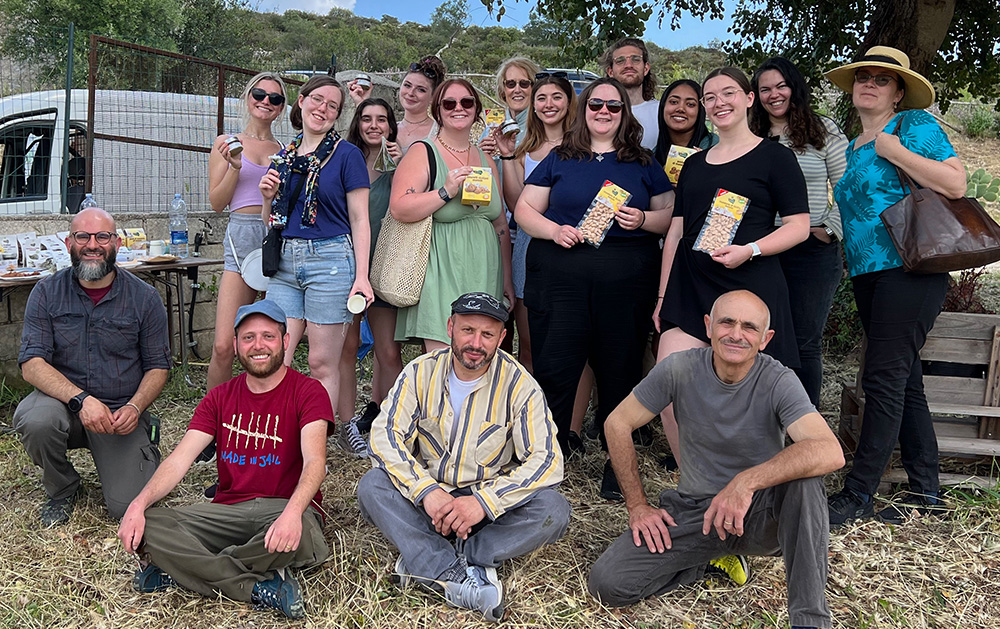
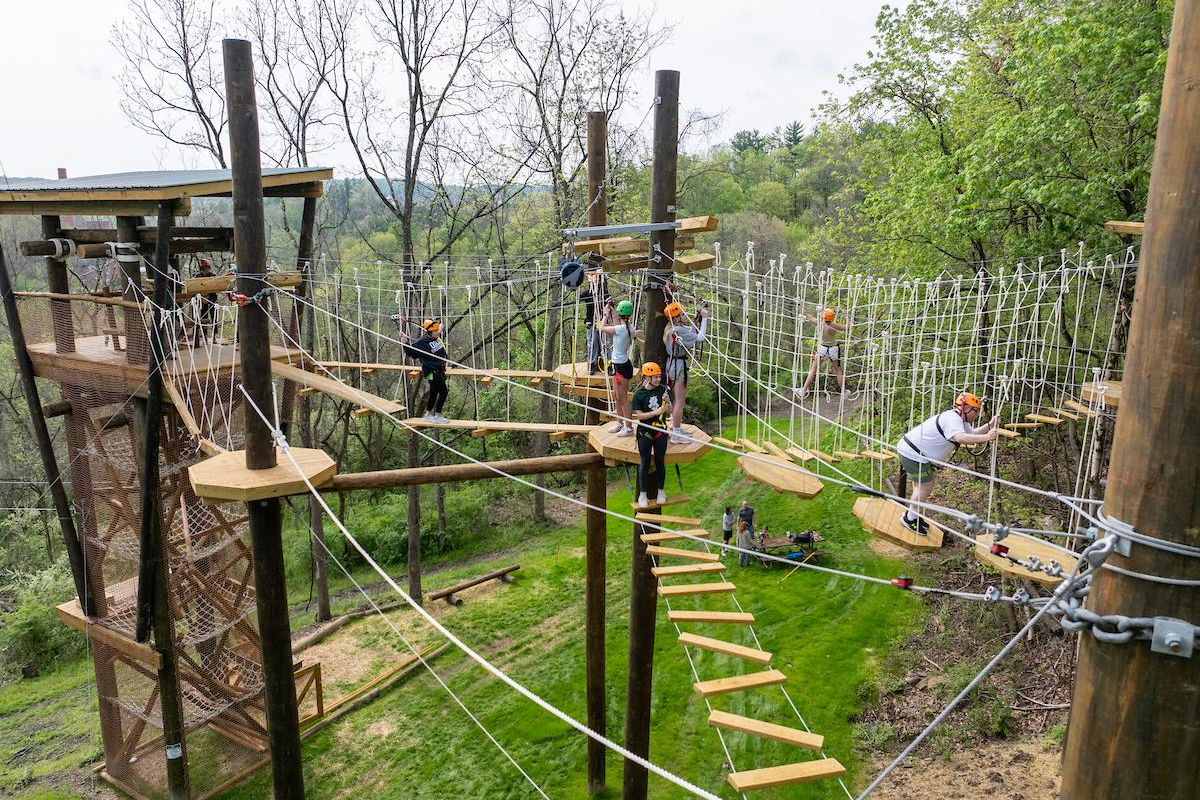
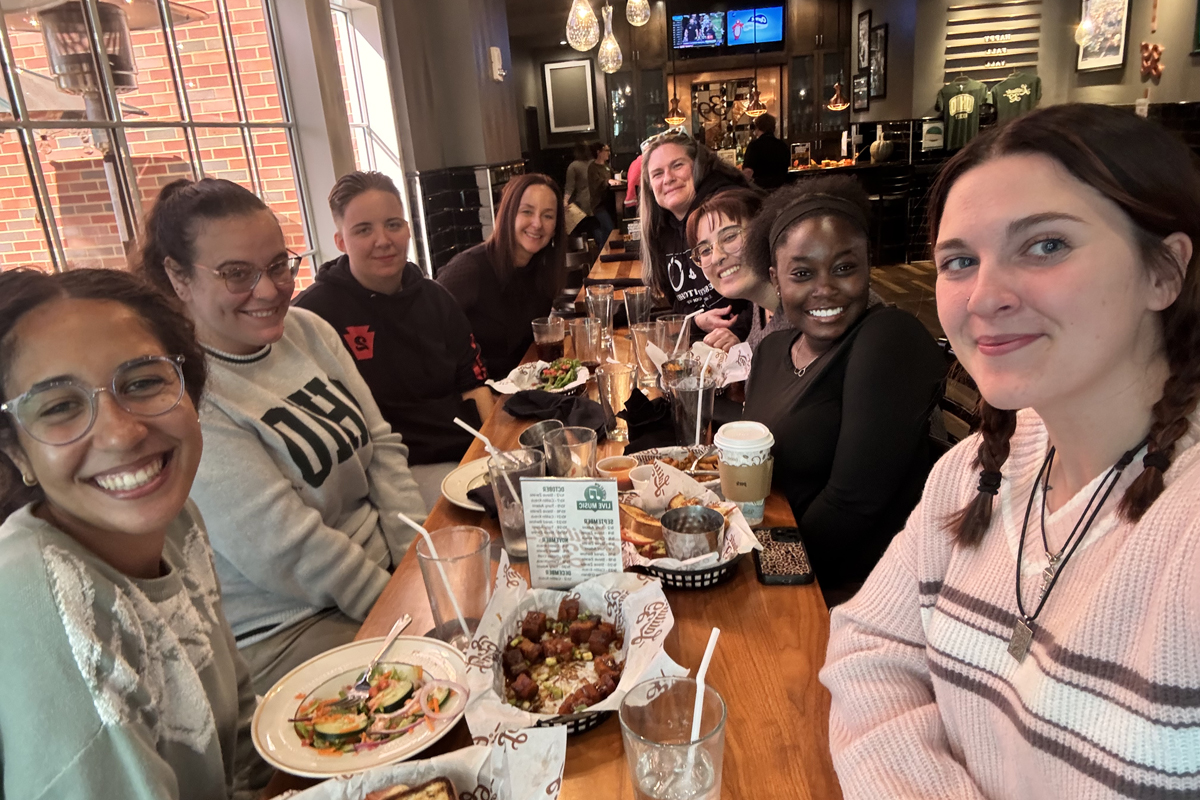
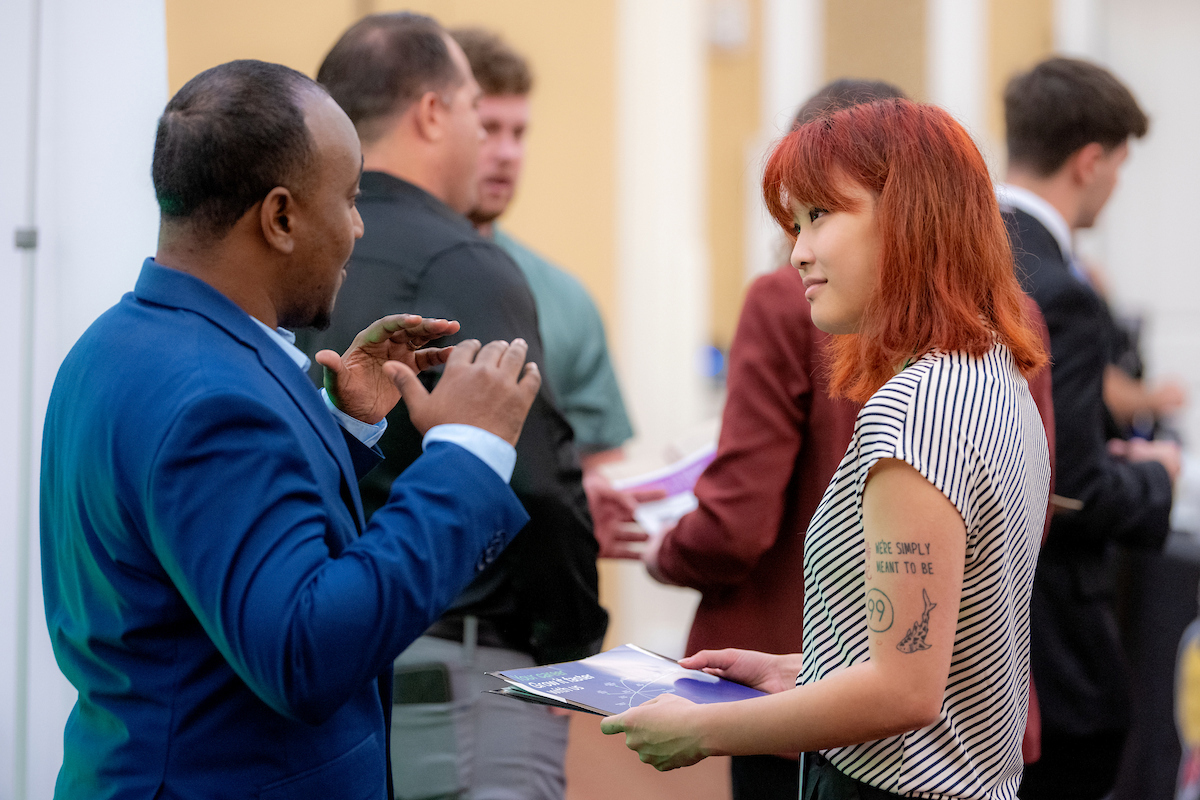
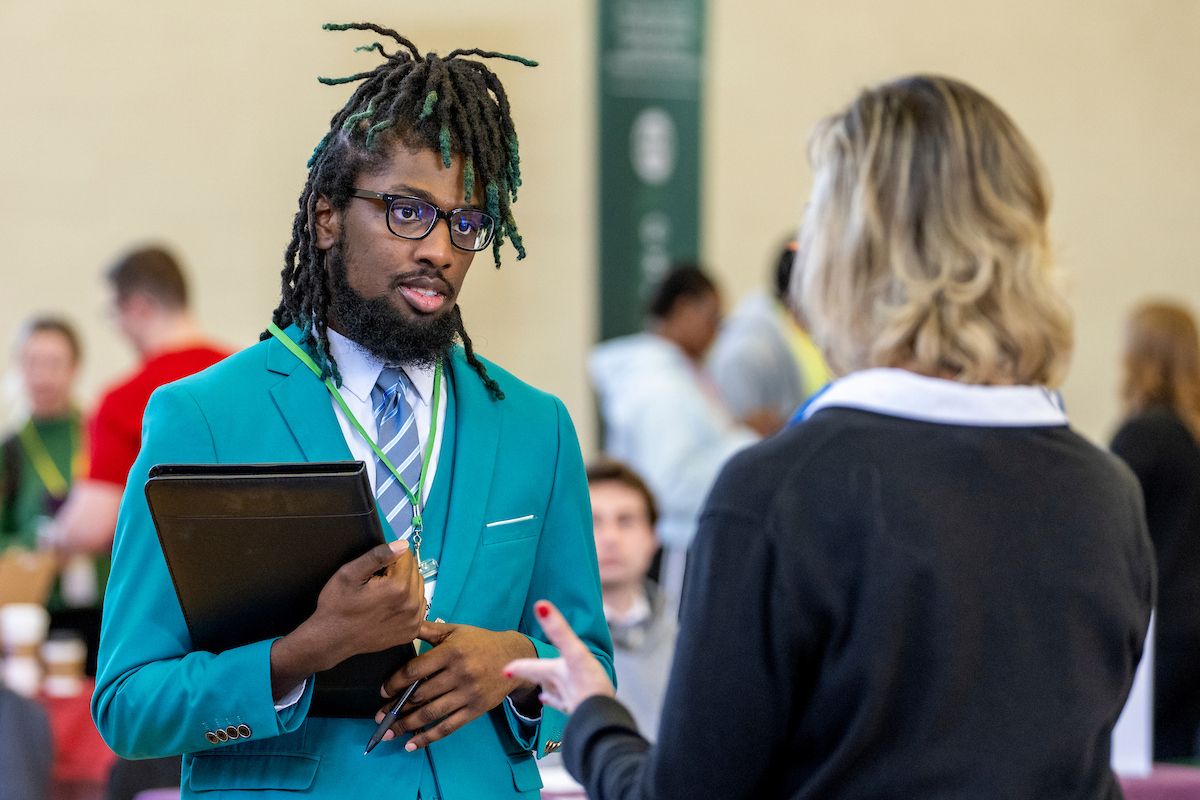
Alumni involvement drives innovation and impact in the College of Arts and Sciences for both its students and faculty. Explore the various areas of opportunity to pair alumni engagement and personal philanthropy with the highest areas of need for our college, including supporting students, experiential learning and investing in faculty.